What are the key differences between KNN and K-means?
Answer
KNN(K-Nearest Neighbors) is a supervised algorithm that classifies data by considering the labels of its nearest neighbors, emphasizing prediction based on historical data. In contrast, K-Means is an unsupervised clustering technique that groups data together based solely on their similarity, without using any labels.
Here are the key differences between KNN and K-Means:
(1) Learning Type
KNN: Supervised learning algorithm (used for classification/regression).
K-Means: Unsupervised learning algorithm (used for clustering).
(2) Objective
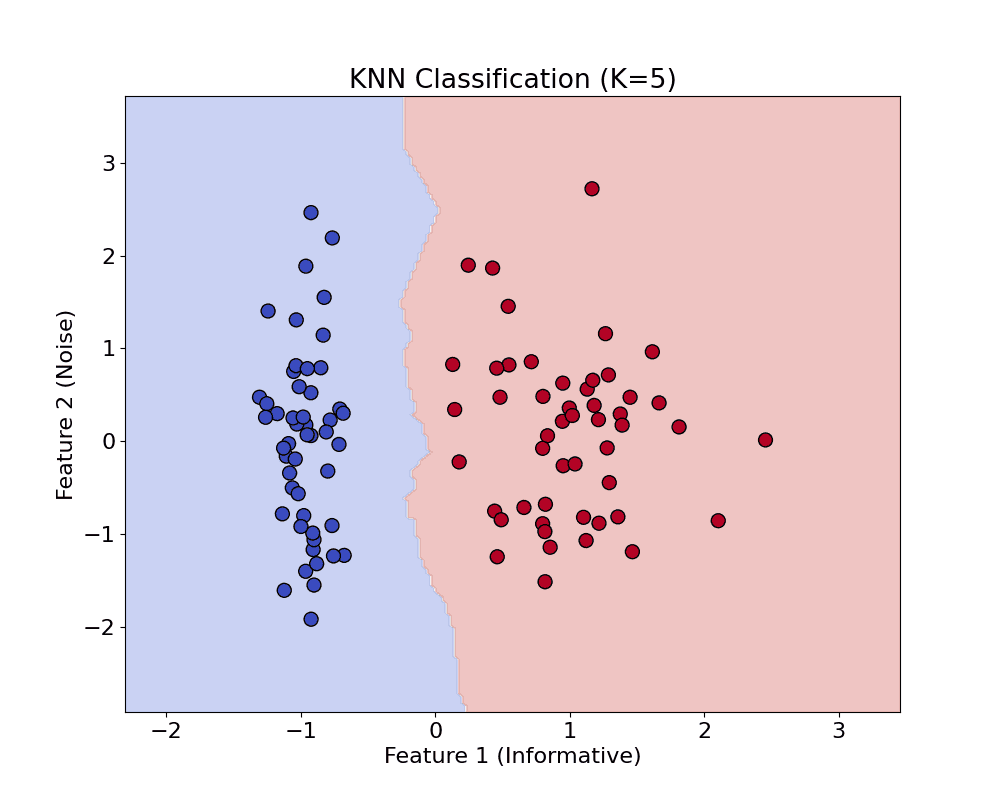
KNN: Predict the label of a new sample based on the majority vote (or average) of its K nearest neighbors.
K-Means: Partition the dataset into K clusters by minimizing intra-cluster distance.
(3) Training
KNN: No explicit training; It simply stores the entire training dataset.
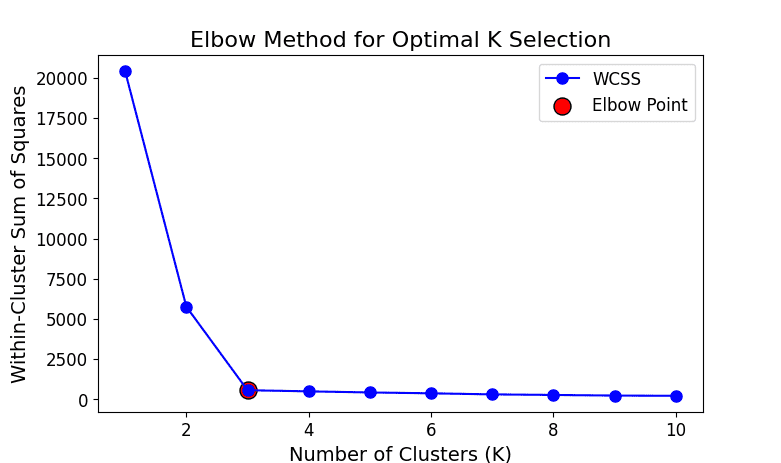
K-Means: Involves an iterative training process to learn cluster centroids.
(4) Prediction
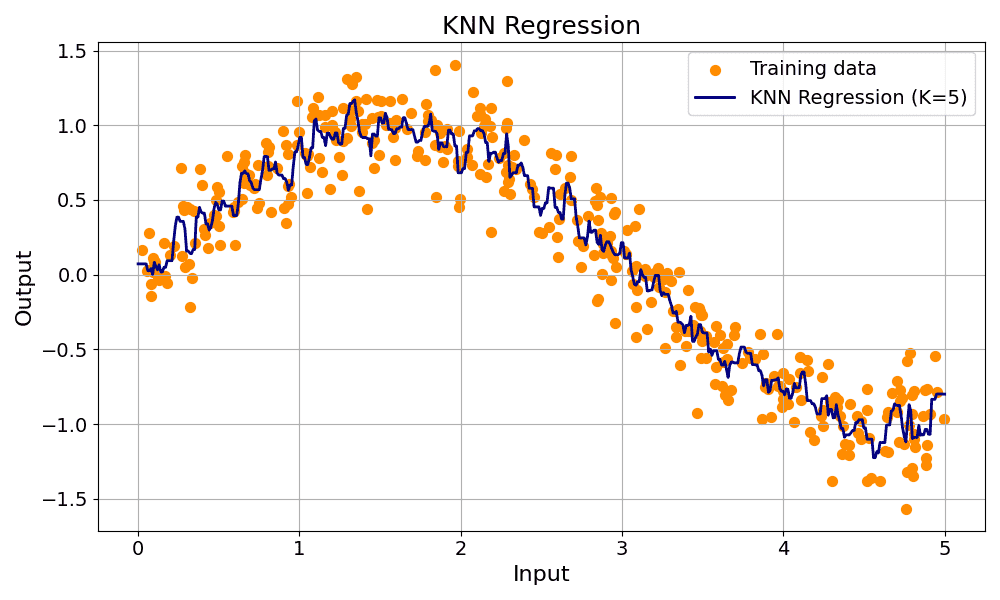
KNN: Computationally expensive, computes the distance from the test point to every training point. Sorts the distances and selects the top nearest neighbors. The majority votes for classification. Average of values for regression.
K-Means: Fast and simple for inference, compute the distance of any new data point to each of the centroids. Assign it to the nearest centroid (i.e., predicted cluster).
(5) Distance Metric Use
KNN: Used to find neighbors.
K-Means: Used to assign points to the nearest cluster center.
(6) Output
KNN: Outputs a label (classification) or value (regression).
K-Means: Outputs cluster assignments and centroids.
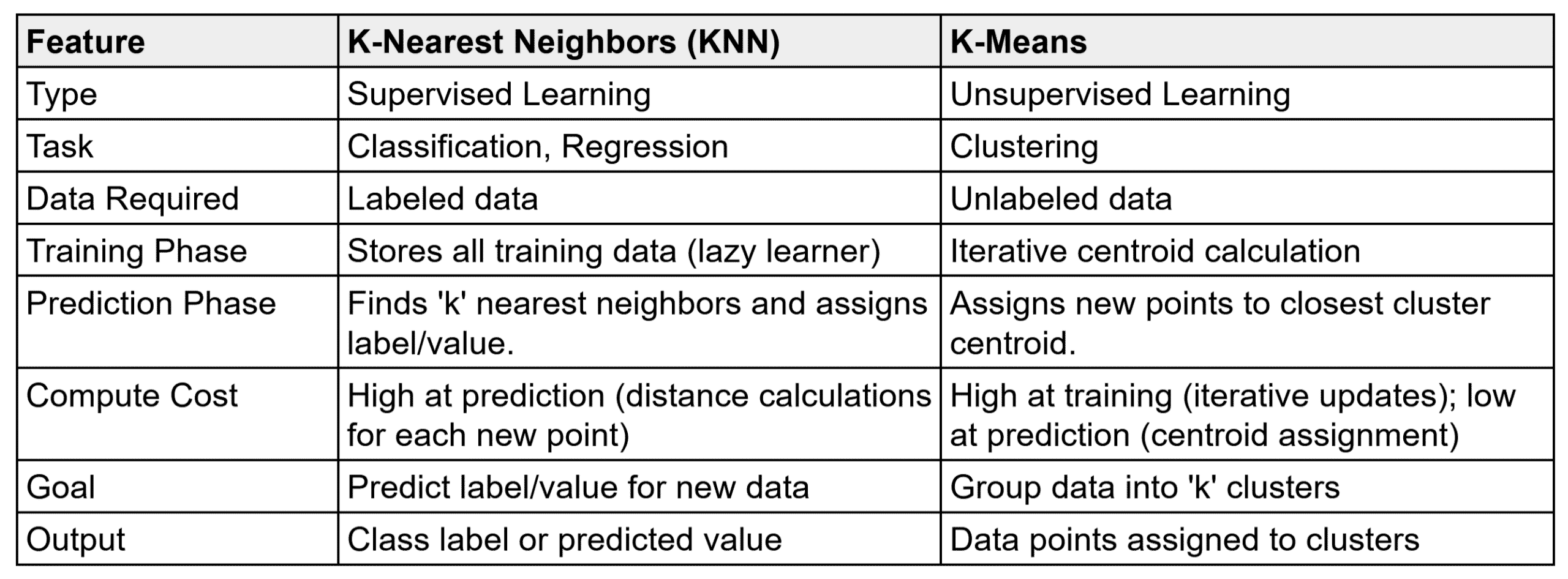
The table below summarizes the comparison between KNN and K-Means.